As in recent years, and certainly in the future, in vitro cell-based experiments are highly relevant in fundamental biological research.
Cell culture in vitro offers the advantage of providing a controlled physicochemical culture environment without systemic variables for the cells. Thus, it can be used to analyze the biological responses of cells to a variety of external stimuli (e.g., changes in temperature or pH, drugs, cytokines, etc.).
Together with analyzing phenotypic changes in cells during biological experiments, information about changes in the transcriptome of treated cells is also critical. This is because not all stimuli can cause morphological changes, however, they can still affect the transcriptional activity of a wide range of genes that govern crucial cellular pathways.
Despite the widespread use of microscopy-based readouts for identifying cellular changes in response to various stimuli, transcriptional analysis is required to clearly define the underlying molecular dynamic in cells. Therefore, gene expression analysis along with phenotypic analysis is significant.
I.DOT- A premier low-volume liquid handler
An asset for high-throughput cell-based assays and gene expression analysis.

Traditional workflows for phenotypic cell-based screening and gene expression analysis of cells in microtiter plates. Which involve multiple steps and larger reagent volumes, which add time, labor, and cost. Furthermore, the multiple-step protocols and the need to transfer the cells to different wells and microtubes throughout the entire experimental procedure increase the loss of input materials, which is particularly critical when working with low cell numbers or single cells.
Moreover, in vitro studies are prone to inter and intra-experimental variation due to consumable properties and human experimental errors.
Therefore, miniaturization and parallelization of experimental steps for sample preparation are useful tools to increase efficiency and reduce the experimental budget and time.
I.DOT, an asset for any laboratory
I.DOT Liquid Handler: a low volume liquid dispenser with the capability to dispense as little as 8 nL up to microliter volumes with unrivaled speed, precision, and accuracy opens a plethora of possibilities for miniaturization of experiment workflows and even enables cell-based assays and gene expression analysis on chip formats.
The I.DOT has proven to be a valuable asset for the authors of the following publication highlighted in this blog. They used the I.DOT for establishing the first miniaturized on-chip strategy (“Cells-to-cDNA on Chip”) that enables the entire course of cell screening, phenotypic microscopy-based analysis. As well as mRNA isolation, and conversion to cDNA for gene expression analysis by real-time PCR on an open DMA platform (Fig: 1).
The authors believe this could be a game changer in high throughput phenotypic analysis and to delineate the underlying molecular dynamics of the cells.
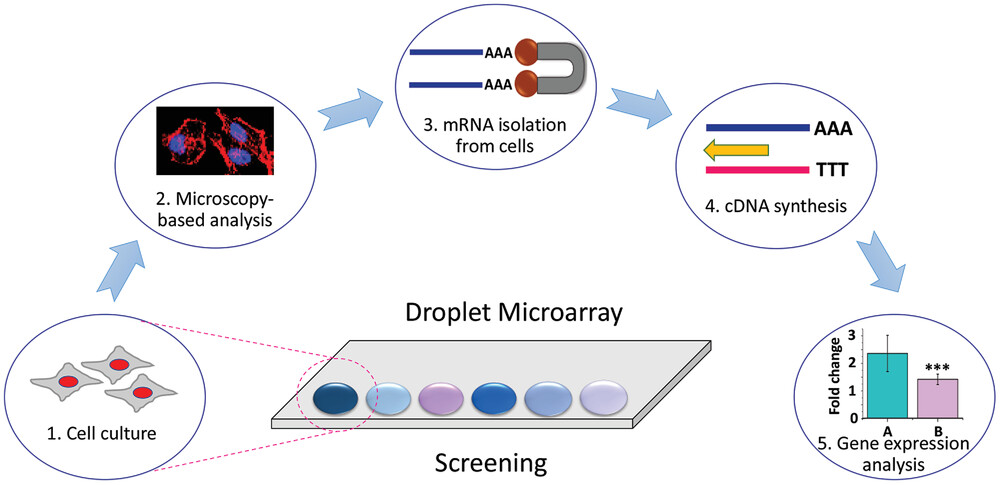
Figure 1*: *Schematic representation of “Cells-to-cDNA on Chip” method. 1. cell culture and screening in micro/nanoliter droplets on DMA, 2. phenotypic analysis of cells by microscopy-based methods, 3. cell lysis and isolation of mRNA using poly-T magnetic beads, 4. mRNA to cDNA conversion within the same droplet, 5. collection of cDNA for qualitative and quantitative gene expression analysis.
*Figure Adapted from https://doi.org/10.1002/adhm.202102493
The I.DOT has been effectively implemented to dispense HeLa-CCL2 cells for cell seeding as well as all reagents required for on-chip mRNA isolation, annealing primer to template RNA, and reverse transcription in volumes as small as 10 nanolitres, thus reducing the reaction volumes up to 100-fold compared to usual lab-based protocol, thereby leading to a drastic reduction in cost, time, and labor. Furthermore, it has eliminated the risk of cross-contamination and produced reproducible and reliable results.
Access the free research article to find out more about the workflow and implementation of I.DOT Liquid Handler
Download: https://doi.org/10.1002/adhm.202102493